Section 6.2: Estimating the Hazard Function
Example 6.1 using data set bone_marrow. The first two steps are to build up an event data set which contains information on the number of events and number of subjects at risk at each time point. Table 6.1 is created based on it with all the calculation on the weight variables.
proc sql;
create table bone_t as
select t2, sum(dfree) as num_event, count(t2) as sub_total
from bone_marrow
where g = 1
group by t2;
quit;
data table6_1a;
set bone_t nobs = all;
total1 = lag(sub_total);
retain num_left 38;
if _n_ > 1 then do;
num_left = - total1 + num_left; end;
retain h sigma2 0;
h = h + num_event / num_left;
sigma2 = sigma2 + num_event /(num_left)**2;
keep t2 num_event num_left h sigma2;
if num_event~=0 or _n_ = all;
run;
%let a_50 = 1.323;
%let b_50 = 1.102;
%let a_600 = 1.148;
%let b_600 = .560;
data table6_1;
set table6_1a nobs=last;
lagh=lag(h);
lagvar=lag(sigma2);
if _n_ = 1 then do;
deltah = h;
deltav = sigma2;
end;
else do;
deltah = h - lagh;
deltav = sigma2 - lagvar;
end;
trans1 = (150 - t2)/100; /*t=150*/
if abs(trans1)> 1 then k1 = 0 ;
else k1 = .75*(1-trans1**2);
trans2 = ( 50 - t2)/100; /*t=50*/
if abs(trans2) > 1 then k2 = 0;
else k2 = .75 *(1-trans2**2)*(&a_50 + &b_50 *trans2);
trans3 = (600 - t2)/100; /*t=600*/
if t2 > 600 then trans3 = - trans3;
if abs(trans3) > 1 then k3 = 0;
else k3 = .75*(1-trans3**2)*(&a_600 + &b_600*trans3);
if _n_ ~= last;
run;
proc print data=table6_1 noobs;
var t2 h deltah deltav trans1 k1 trans2 k2 trans3 k3;
run;
t2 h deltah deltav trans1 k1 trans2 k2 trans3 k3 1 0.02632 0.026316 .000692521 1.49 0.00000 0.49 1.06176 5.99 0.00000 55 0.05334 0.027027 .000730460 0.95 0.07313 -0.05 0.94855 5.45 0.00000 74 0.08112 0.027778 .000771605 0.76 0.31680 -0.24 0.74816 5.26 0.00000 86 0.10969 0.028571 .000816327 0.64 0.44280 -0.36 0.60468 5.14 0.00000 104 0.13910 0.029412 .000865052 0.46 0.59130 -0.54 0.38674 4.96 0.00000 107 0.16941 0.030303 .000918274 0.43 0.61133 -0.57 0.35182 4.93 0.00000 109 0.20066 0.031250 .000976563 0.41 0.62393 -0.59 0.32896 4.91 0.00000 110 0.23291 0.032258 .001040583 0.40 0.63000 -0.60 0.31766 4.90 0.00000 122 0.29958 0.066667 .002222222 0.28 0.69120 -0.72 0.19128 4.78 0.00000 129 0.33530 0.035714 .001275510 0.21 0.71693 -0.79 0.12755 4.71 0.00000 172 0.37233 0.037037 .001371742 -0.22 0.71370 -1.22 0.00000 4.28 0.00000 192 0.41079 0.038462 .001479290 -0.42 0.61770 -1.42 0.00000 4.08 0.00000 194 0.45079 0.040000 .001600000 -0.44 0.60480 -1.44 0.00000 4.06 0.00000 230 0.49427 0.043478 .001890359 -0.80 0.27000 -1.80 0.00000 3.70 0.00000 276 0.53973 0.045455 .002066116 -1.26 0.00000 -2.26 0.00000 3.24 0.00000 332 0.58735 0.047619 .002267574 -1.82 0.00000 -2.82 0.00000 2.68 0.00000 383 0.63735 0.050000 .002500000 -2.33 0.00000 -3.33 0.00000 2.17 0.00000 418 0.68998 0.052632 .002770083 -2.68 0.00000 -3.68 0.00000 1.82 0.00000 466 0.74553 0.055556 .003086420 -3.16 0.00000 -4.16 0.00000 1.34 0.00000 487 0.80436 0.058824 .003460208 -3.37 0.00000 -4.37 0.00000 1.13 0.00000 526 0.86686 0.062500 .003906250 -3.76 0.00000 -4.76 0.00000 0.74 0.53012 609 0.93829 0.071429 .005102041 -4.59 0.00000 -5.59 0.00000 0.09 0.89152 662 1.01521 0.076923 .005917160 -5.12 0.00000 -6.12 0.00000 0.62 0.69033
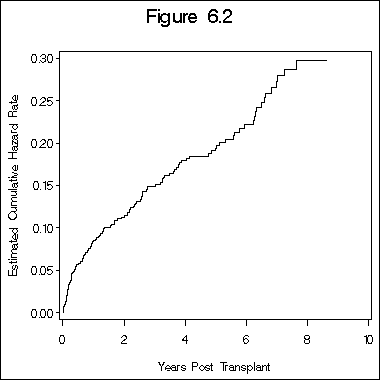
Example 6.2 on the kidney transplant data. You can download the data step that creates the dataset called sec1_7 here.
data fig6_1; set sec1_7; timey = time/365.5; run; proc phreg data = fig6_1; model timey*death(0) = ; output out = fig6_2 logsurv=h; run; data fig6_2a; set fig6_2; haz = - h; run; proc sort data=fig6_2a; by timey; symbol i = stepjl v=none; axis1 order = (0.0 to .30 by .05) minor = none label=(a=90); axis2 order = (0 to 10 by 2) minor = none; title "Figure 6.2"; proc gplot data = fig6_2a; plot haz*timey /vaxis = axis1 haxis=axis2; label haz = "Estimated Cumulative Hazard Rate"; label timey = "Years Post Transplant"; run; quit;
Section 6.3: Estimation of Excess Mortality
This section uses the 1959-1960 Iowa State life tables. We created a SAS data step for it. The data set for this section is the data on the 26 psychiatric patients in Iowa described in section 1.15.
Example 6.3, Table 6.3 and Figure 6.8. The first part of the program creates the hazard rate based on the Iowa life table. The macro does the computation for table 6.3 including the confidence intervals. We notice that the output produced below does not match with Table 6.3 completely. The data for table 6.3 is then used for Figure 6.8.
proc sort data = iowa_60_mortality;
by male;
run;
data table6_2;
set iowa_60_mortality;
by male;
ls = lag(survival);
if ~first.male then do;
hr = log(ls) - log(survival); end;
drop survival;
if hr ~=.;
run;
%macro rel_mortality(data, age, time, death, outdata);
proc sql noprint;
select count( distinct &time) into :max_num
from &data
where &death = 1;
quit;
proc sql noprint;
select count(&time) into :dpoint separated by ' '
from &data
where &death = 1
group by &time;
quit;
proc sql noprint;
select distinct &time into :tpoint separated by ' '
from &data
where &death = 1;
quit;
%do k = 1 %to &max_num;
%let p = %scan(&tpoint, &k);
proc sql noprint;
select sum(y.hr) into :q&k
from &data as x, table6_2 as y
where 1-x.female = y.male and x.age + &p + 1 = y.age and x.time >= &p ;
quit;
%end;
data &outdata;
b = 0;
v = 0;
%do i = 1 %to &max_num;
t = %scan(&tpoint, &i);
d = %scan(&dpoint, &i);
q = &&q&i;
b = b + d/q;
v = v + d/q**2;
s = sqrt(v);
lc = b/exp(1.96*s/b);/*log-transformed confidence interval*/
uc = b*exp(1.96*s/b);
output;
%end;
run;
proc print data= &outdata noobs;
var t d q b v s lc uc;
run;
%mend rel_mortality;
%rel_mortality(sec1_15, age, time, death, temp)
t d q b v s lc uc
1 2 0.05932 33.718 568.44 23.8420 8.4325 134.822
2 1 0.04964 53.861 974.20 31.2122 17.2982 167.707
11 1 0.08524 65.593 1111.84 33.3443 24.2183 177.654
14 1 0.10278 75.323 1206.51 34.7349 30.5066 185.979
22 2 0.18238 86.289 1266.64 35.5899 38.4478 193.660
24 1 0.21559 90.928 1288.15 35.8909 41.9472 197.100
25 1 0.17002 96.809 1322.75 36.3696 46.3583 202.164
26 1 0.19441 101.953 1349.20 36.7315 50.3179 206.574
28 1 0.18434 107.377 1378.63 37.1299 54.5220 211.473
32 1 0.18562 112.765 1407.66 37.5187 58.7436 216.465
35 1 0.16755 118.733 1443.28 37.9905 63.4180 222.296
40 1 0.04902 139.135 1859.50 43.1219 75.7912 255.419
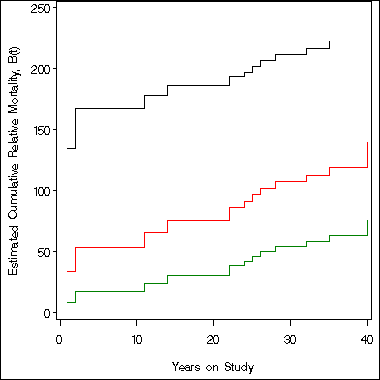
symbol i = stepjl;
axis1 order = (0 to 250 by 50) label=(a=90 'Estimated Cumulative Relative Mortality, B(t)')
minor = none;
axis2 order = (0 to 40 by 10) minor = none label=('Years on Study');
proc gplot data= temp;
plot uc*t b*t lc*t/overlay vaxis = axis1 haxis = axis2;
run;
quit;
Example 6.3 continued on page 169. Table 6.4, Figure 6.9 and Figure 6.10. The proc sql step and data steps below prepares us to create Table 6.4. The macro program generates Table 6.4.
proc sql;
create table table6_4 as
select time, sum(death) as d, sum(death=0) as censor
from sec1_15
group by time;
quit;
data table6_4a;
do time = 1 to 40;
output; end;
run;
data table6_4b;
merge table6_4 table6_4a ;
by time;
if d = . then d = 0;
if censor = . then censor = 0;
retain y 26;
ld = lag(d);
lc = lag(censor);
if _n_>1 then y = y-ld - lc;
drop ld lc;
run;
%macro theta(data, time, max, outdata);
%do k = 1 %to &max;
proc sql noprint;
select sum(y.hr) into :q&k
from &data as x, table6_2 as y
where 1-x.female = y.male and x.age + &k = y.age and x.time > &k - 1 ;
quit;
%end;
data &outdata;
set table6_4b;
retain theta 0;
retain p var_a 0;
retain h 0;
retain s1 1;
q = symget('q'||left(_n_));
theta = theta + q/y;
p = p + d/y;
h = h + d/y;
s1 = s1*(1-d/y);
var_a = var_a + d/y**2;
a = p - theta;
sda = sqrt(var_a);
survival = exp(-theta);
sc = s1/survival;
drop q p censor;
run;
proc print data = temp noobs;
var time d y h theta a sda s1 survival sc;
run;
%mend;
%theta(sec1_15, time, 40, temp)
time d y h theta a sda s1 survival sc
1 2 26 0.07692 0.00213 0.07479 0.05439 0.92308 0.99787 0.92505
2 1 24 0.11859 0.00406 0.11453 0.06852 0.88462 0.99595 0.88822
3 0 23 0.11859 0.00594 0.11265 0.06852 0.88462 0.99408 0.88988
4 0 23 0.11859 0.00794 0.11065 0.06852 0.88462 0.99209 0.89167
5 0 23 0.11859 0.01008 0.10851 0.06852 0.88462 0.98997 0.89358
6 0 23 0.11859 0.01237 0.10622 0.06852 0.88462 0.98771 0.89563
7 0 23 0.11859 0.01483 0.10376 0.06852 0.88462 0.98528 0.89783
8 0 23 0.11859 0.01747 0.10112 0.06852 0.88462 0.98269 0.90020
9 0 23 0.11859 0.02032 0.09827 0.06852 0.88462 0.97989 0.90277
10 0 23 0.11859 0.02341 0.09518 0.06852 0.88462 0.97686 0.90557
11 1 23 0.16207 0.02679 0.13528 0.08115 0.84615 0.97357 0.86913
12 0 22 0.16207 0.03029 0.13178 0.08115 0.84615 0.97016 0.87218
13 0 22 0.16207 0.03414 0.12793 0.08115 0.84615 0.96643 0.87554
14 1 22 0.20752 0.03838 0.16914 0.09301 0.80769 0.96235 0.83929
15 0 21 0.20752 0.04279 0.16473 0.09301 0.80769 0.95811 0.84300
16 0 21 0.20752 0.04811 0.15941 0.09301 0.80769 0.95303 0.84750
17 0 21 0.20752 0.05297 0.15455 0.09301 0.80769 0.94841 0.85163
18 0 21 0.20752 0.05882 0.14871 0.09301 0.80769 0.94288 0.85662
19 0 21 0.20752 0.06522 0.14230 0.09301 0.80769 0.93686 0.86213
20 0 21 0.20752 0.07222 0.13530 0.09301 0.80769 0.93032 0.86818
21 0 21 0.20752 0.08035 0.12717 0.09301 0.80769 0.92280 0.87527
22 2 21 0.30276 0.08872 0.21405 0.11483 0.73077 0.91511 0.79856
23 0 19 0.30276 0.09674 0.20603 0.11483 0.73077 0.90780 0.80499
24 1 19 0.35539 0.10611 0.24928 0.12632 0.69231 0.89932 0.76981
25 1 18 0.41095 0.11683 0.29411 0.13800 0.65385 0.88973 0.73488
26 1 17 0.46977 0.12554 0.34423 0.15001 0.61538 0.88202 0.69770
27 0 16 0.46977 0.13628 0.33349 0.15001 0.61538 0.87260 0.70524
28 1 16 0.53227 0.14737 0.38490 0.16251 0.57692 0.86297 0.66853
29 0 15 0.53227 0.15924 0.37303 0.16251 0.57692 0.85279 0.67651
30 0 15 0.53227 0.17294 0.35933 0.16251 0.57692 0.84119 0.68584
31 0 13 0.53227 0.18722 0.34505 0.16251 0.57692 0.82926 0.69571
32 1 11 0.62318 0.20356 0.41962 0.18621 0.52448 0.81582 0.64288
33 0 10 0.62318 0.22147 0.40171 0.18621 0.52448 0.80134 0.65450
34 0 8 0.62318 0.24193 0.38125 0.18621 0.52448 0.78511 0.66803
35 1 7 0.76604 0.26380 0.50223 0.23470 0.44955 0.76812 0.58526
36 0 4 0.76604 0.28556 0.48048 0.23470 0.44955 0.75160 0.59813
37 0 3 0.76604 0.31402 0.45202 0.23470 0.44955 0.73051 0.61540
38 0 2 0.76604 0.35176 0.41428 0.23470 0.44955 0.70345 0.63907
39 0 2 0.76604 0.39333 0.37271 0.23470 0.44955 0.67481 0.66619
40 1 1 1.76604 0.43709 1.32895 1.02717 0.00000 0.64591 0.00000
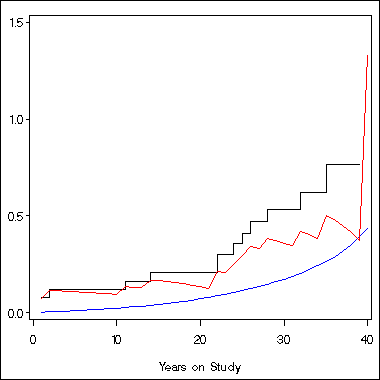
axis1 order = (0 to 1.5 by .5) label=(' ') minor = none;
axis2 order = (0 to 40 by 10) label=('Years on Study') minor = none;
symbol1 i = stepjl c = black;
symbol2 i = join c = blue;
symbol3 i = join c = red;
proc gplot data = temp;
plot h*time = 1 theta*time = 2 a*time = 3 /overlay vaxis = axis1 haxis=axis2;
run;
quit;
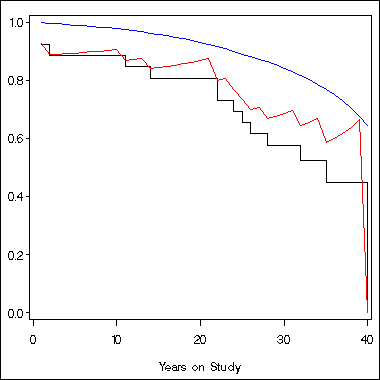
axis1 order = (0 to 1 by .2) label=(' ') minor = none;
axis2 order = (0 to 40 by 10) label=('Years on Study') minor = none;
symbol1 i = stepjl c = black;
symbol2 i = join c = blue;
symbol3 i = join c = red;
proc gplot data = temp;
plot s1*time = 1 survival*time = 2 sc*time = 3 /overlay vaxis = axis1
haxis=axis2;
run;
quit;
Section 6.4: Bayesian Nonparametric Methods
Under construction.