In this chapter we will be using the uis data sets.
Creating the interactions needed for the model specified in table 5.11, p. 213.
data uis; set uis; ivhx3 = (ivhx = 3); ndrugfp1 = 1/((ndrugtx+1)/10); ndrugfp2 = (1/((ndrugtx+1)/10))*log((ndrugtx+1)/10); racesite = race*site; agesite = age*site; run;
Table 6.4, p. 213.
Testing the proportionality assumptions by including the interactions of all the main effects with log(time). The tests of the parameter estimates are tests of the individual interaction. Note that none of which are significant at the 0.05 level.
proc phreg data=uis;
model time*censor(0) = age becktota ndrugfp1 ndrugfp2 ivhx3 race treat site
racesite agesite aget beckt fp1t ivhx3t racet treatt sitet;
aget = age*log(time);
fp1t = ndrugfp1*log(time);
fp2t = ndrugfp2*log(time);
beckt = becktota*log(time);
ivhx3t = ivhx3*log(time);
racet = race*log(time);
treatt = treat*log(time);
sitet = site*log(time);
run;
<output omitted>
Model Fit Statistics
Without With
Criterion Covariates Covariates
-2 LOG L 5327.970 5255.298
Analysis of Maximum Likelihood Estimates
Parameter Standard Hazard
Variable DF Estimate Error Chi-Square Pr > ChiSq Ratio
aget 1 0.00174 0.00850 0.0419 0.8377 1.002
beckt 1 -0.00712 0.00514 1.9178 0.1661 0.993
fp1t 1 -0.01560 0.01759 0.7864 0.3752 0.985
ivhx3t 1 -0.03007 0.11334 0.0704 0.7907 0.970
racet 1 0.11344 0.12497 0.8240 0.3640 1.120
treatt 1 0.12759 0.09978 1.6350 0.2010 1.136
sitet 1 -0.02264 0.11370 0.0397 0.8422 0.978
Comparing the model with the interactions with log(time) to the model without those interactions. The test statistic G = 5260.836 – 5255.298 = 5.538 which is compared to a chi-squared distribution with 7 degrees of freedom which results in a p-value of .595.
proc phreg data=uis;
model time*censor(0) = age becktota ndrugfp1 ndrugfp2 ivhx3 race treat site racesite agesite;
run;
<output omitted>
Model Fit Statistics
Without With
Criterion Covariates Covariates
-2 LOG L 5327.970 5260.836
Fig. 6.5, p. 214.
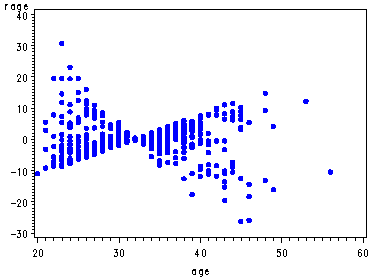
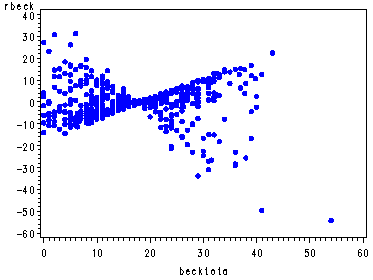
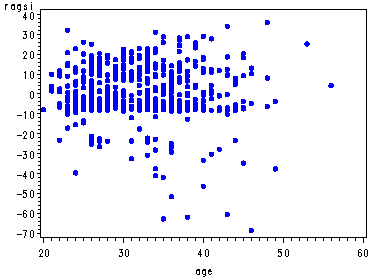
The graphs of the score residuals. Note that there is a score residual for each predictor in the model and that in order to output all of them they each have to be named in a list following the ressco option in the output statement. The order of their names must match the order in which they appear in the model statement.
proc phreg data=uis noprint; model time*censor(0) = age becktota ndrugfp1 ndrugfp2 ivhx3 race treat site racesite agesite; output out=res ressco=rage rbeck rfp1 rfp2 riv rrace rtreat rsite rrasi ragsi; run; goptions reset=all; symbol v=dot h=.8 c=blue; proc gplot data=res; plot rage*age; plot rbeck*becktota; plot ragsi*age; run; quit;
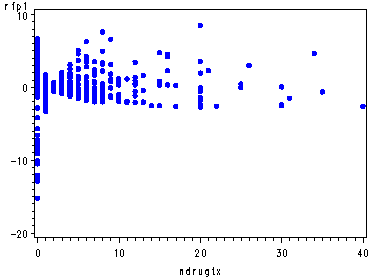
Since ndrugtx was not a predictor in the model and therefore it is not included in the outputted data set. So, we must merge the outputted data set containing the score residuals with the original dataset containing the predictor ndrugtx. Then, finally, we will be able to plot.
data residual; set res; keep time rfp1; run; proc sort data=residual; by time; run; data ndrugtx; set uis; keep time ndrugtx; run; proc sort data=ndrugtx; by time; run; data residdrug; merge residual ndrugtx; by time; run; goptions reset=all; symbol v=dot h=.8 c=blue; proc gplot data=residdrug; plot rfp1*ndrugtx; run; quit;
Table 6.6, p. 230.
The covout option with the outest option outputs a dataset containing the variance/covariance matrix. These numbers are used in the calculations on p. 233-234 and p. 236-237, where the 95% interval for the estimate of specific covariate patterns are being calculated.
proc phreg data=uis covout outest=covariance;
model time*censor(0) = age becktota ndrugfp1 ndrugfp2 ivhx3 race treat site racesite agesite;
run;
proc print data=covariance;
where _name_ ne "time";
var _name_ age becktota ndrugfp1 ndrugfp2 ivhx3 race treat site racesite agesite;
run;
<output omitted>
Analysis of Maximum Likelihood Estimates
Parameter Standard Hazard
Variable DF Estimate Error Chi-Square Pr > ChiSq Ratio
age 1 -0.04140 0.00991 17.4395 <.0001 0.959
becktota 1 0.00874 0.00497 3.0968 0.0784 1.009
ndrugfp1 1 -0.57446 0.12519 21.0567 <.0001 0.563
ndrugfp2 1 -0.21458 0.04859 19.5043 <.0001 0.807
ivhx3 1 0.22775 0.10856 4.4009 0.0359 1.256
race 1 -0.46689 0.13476 12.0039 0.0005 0.627
_NAME_ age becktota ndrugfp1 ndrugfp2 ivhx3
age 0.000098263 0.000002698 0.000266 0.000094871 -0.000196
becktota 0.000002698 0.000024655 -0.000036 -.000015147 -0.000059
ndrugfp1 0.000265660 -.000036449 0.015672 0.006022342 0.001566
ndrugfp2 0.000094871 -.000015147 0.006022 0.002360698 0.000437
ivhx3 -.000195980 -.000058749 0.001566 0.000436894 0.011786
race 0.000003095 -.000037065 0.000464 0.000215281 0.003110
treat 0.000087378 0.000008921 0.000079 0.000009281 0.000663
site 0.002863893 0.000118416 0.008445 0.003553242 0.004147
racesite -.000089174 0.000001051 -0.001732 -.000583900 -0.002285
agesite -.000089888 -.000003222 -0.000223 -.000092924 -0.000031
race treat site racesite agesite
0.000003 0.000087378 0.00286 -0.000089 -.000089888
-0.000037 0.000008921 0.00012 0.000001 -.000003222
0.000464 0.000079096 0.00845 -0.001732 -.000222889
0.000215 0.000009281 0.00355 -0.000584 -.000092924
0.003110 0.000663059 0.00415 -0.002285 -.000031018
0.018159 -.000044335 0.00805 -0.017717 -.000091465
-0.000044 0.008900028 0.00637 -0.000930 -.000172920
0.008045 0.006373764 0.28243 -0.006604 -.008320538
-0.017717 -.000929914 -0.00660 0.061384 -.000236448
-0.000091 -.000172920 -0.00832 -0.000236 0.000258588