Table 13.1, page 471
Sorting the data to get it in the same order as in the table in the book.
proc sort data=datasets.honking out=sort_honk; by seconds; run; proc print data=sort_honk; id id; var seconds censor; run; ID SECONDS CENSOR 9 1.41 0 54 1.41 1 40 1.51 0 50 1.67 0 60 1.68 0 7 1.86 0 17 2.12 0 31 2.19 0 3 2.36 1 56 2.48 0 5 2.50 0 22 2.53 0 23 2.54 0 45 2.56 0 27 2.62 0 4 2.68 0 49 2.76 1 21 2.78 1 13 2.83 0 ...
-----------------------------------------------------------------------------------------
Table 13.2, page 477
Life table for the horn-honking data with discrete-time and actuarial estimates of the survivor functions. First, we will produce the survivor and hazard functions using actuarial estimates.
The option method allows us to specify that we wish to obtain the actuarial estimates.
ods output LifetableEstimates=act;
proc lifetest data=datasets.honking method=lt intervals=1 to 8 by 1, 18;
time seconds*censor(1);
run;
proc print data=act noobs;
var lowertime uppertime failed censored survival stderr hazard hazstderr;
run;
<output omitted>
Lower Upper HazStd
Time Time Failed Censored Survival StdErr Hazard Err
0 1 0 0 1.0000 0 0 .
1 2 5 1 1.0000 0 0.092593 0.041364
2 3 14 3 0.9115 0.0378 0.329412 0.086837
3 4 9 2 0.6537 0.0643 0.315789 0.103943
4 5 6 4 0.4754 0.0690 0.333333 0.134179
5 6 2 2 0.3396 0.0680 0.181818 0.128033
6 7 2 2 0.2830 0.0674 0.285714 0.199958
7 8 1 0 0.2122 0.0666 0.222222 0.220846
8 18 3 1 0.1698 0.0654 0.15 0.057282
We now produce the estimates for the discrete-time survivor function.
In the data step we are creating a categorical time variable which is used in the following proc lifetest from which we output the product limit estimates into a data set called est. We remove the redundant observations with missing values for the survival function from the data set and then print.
data honking; set datasets.honking; if seconds < 2 then time = 1; else if seconds < 3 then time = 2; else if seconds < 4 then time = 3; else if seconds < 5 then time = 4; else if seconds < 6 then time = 5; else if seconds < 7 then time = 6; else if seconds < 8 then time = 7; else time = 8; run; ods output ProductLimitEstimates=est; ods listing close; proc lifetest data=honking; time time*censor(1); run; ods listing; data est; set est; if survival; seconds = time + 1; if time=8 then seconds=18; keep survival time seconds stderr; run; proc print data=est noobs; var time survival stderr; run; time Survival StdErr 0.00000 1.0000 0 1.00000 0.9123 0.0375 2.00000 0.6619 0.0632 3.00000 0.4867 0.0683 4.00000 0.3597 0.0673 5.00000 0.3044 0.0674 6.00000 0.2367 0.0673 7.00000 0.1894 0.0685 8.00000 0.0473 0.0444
-----------------------------------------------------------------------------------------
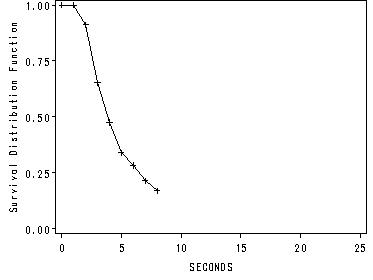
Figure 13.1, page 479
Upper left panel.
We obtain the graph by using the option plots in the proc lifetest statement.
goptions reset=all; proc lifetest data=datasets.honking method=lt intervals=1 to 8 by 1, 18 plots=(s); time seconds*censor(1); run; <output omitted>
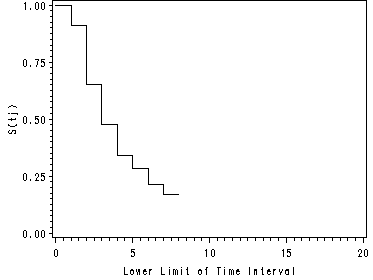
Upper right panel of Figure 13.1
We are using the data set act containing the actuarial estimates.
data act; set act; keep lowertime survival; rename survival = survact; rename lowertime = seconds; run; goptions reset=all; symbol c=blue i=stepjr; axis1 label=(a=90 'S(tj)') order=(0 to 1.00 by 0.25); axis2 order=(0 to 20 by 5); proc gplot data=act; format survact 4.2; plot survact*seconds / overlay vaxis=axis1 haxis=axis2; run; quit;
-----------------------------------------------------------------------------------------
Table 13.3, page 484
We use the ods output statement to output the Kaplan-Meier estimates into a data set called km. Then we drop the observations where survival is missing.
ods output ProductLimitEstimates=km; ods listing close; proc lifetest data=datasets.honking method=km; time seconds*censor(1); run; ods listing; data km; set km; by seconds; if first.seconds; rename survival = survkm; keep seconds survival stderr left; run; proc print data=km noobs; var seconds left survkm stderr; run; SECONDS Left survkm StdErr 0.0000 57 1.0000 0 1.4100 56 0.9825 0.0174 1.5100 54 0.9646 0.0246 1.6700 53 0.9467 0.0299 1.6800 52 0.9289 0.0343 1.8600 51 0.9110 0.0380 2.1200 50 0.8931 0.0412 2.1900 49 0.8753 0.0441 2.4800 47 0.8570 0.0468 2.5000 46 0.8388 0.0492 2.5300 45 0.8206 0.0514 2.5400 44 0.8023 0.0534 2.5600 43 0.7841 0.0552 2.6200 42 0.7659 0.0569 2.6800 41 0.7476 0.0584 2.8300 38 0.7285 0.0599 2.8800 37 0.7093 0.0614 2.8900 36 0.6901 0.0626 2.9200 35 0.6710 0.0637 ... 3.5600 26 0.5325 0.0688 3.5700 25 0.5121 0.0692 ... 4.9600 13 0.3349 0.0683 5.3900 11 0.3070 0.0681 5.7300 10 0.2791 0.0674 6.0300 8 0.2481 0.0666 6.3000 6 0.2126 0.0659 7.2000 4 0.1701 0.0650 9.5900 3 0.1276 0.0611 12.2900 2 0.0851 0.0535 13.1800 1 0.0425 0.0403
-----------------------------------------------------------------------------------------
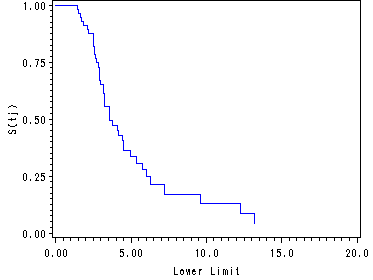
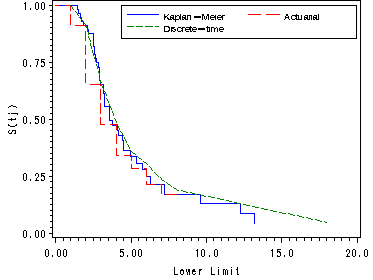
Figure 13.2, page 485
All the data sets containing the survival functions (est, act and km) are sorted and merged in order to graph all the survival functions in one graph.
proc sort data=est;
by seconds;
run;
proc sort data=act;
by seconds;
run;
proc sort data=km;
by seconds;
run;
data combo;
merge km act est;
by seconds;
run;
goptions reset=all;
symbol1 c=blue i=stepjll;
symbol2 c=red i=stepjr line=21;
symbol3 c=green i=j line=3;
axis1 label=(a=90 'S(tj)');
axis2 order=(0 to 20 by 5);
legend label=none value=(height=1 font=swiss 'Kaplan-Meier' 'Actuarial' 'Discrete-time' )
position=(top right inside) mode=share cborder=black;
proc gplot data=combo;
format survkm seconds 4.2;
plot survkm*seconds / vaxis=axis1 haxis=axis2;
plot (survkm survact survival)*seconds / overlay vaxis=axis1 haxis=axis2 legend=legend;
run;
quit;
-----------------------------------------------------------------------------------------
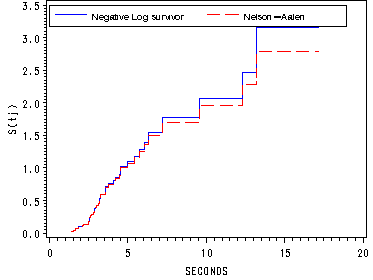
Figure 13.4, page 493
The Nelson-Aalen and negative log survivor function estimates of the cumulative hazard function for the horn-honking dataset.
In the output statement of proc phreg the option method is used to specify the Nelson-Aalen estimates. These estimates are outputted into a data set called nelson which is then sorted and all the redundant observations are dropped.
proc phreg data=datasets.honking noprint; model seconds*censor(1) = ; output out=nelson logsurv=logs / method=emp; run; proc sort data=nelson; by seconds; run; data nelson; set nelson; nlogs = -logs; by seconds; if first.seconds; run;
In this proc phreg we use the output statement to output the product-limit estimates of the log survival function to a data set called pl. The pl data set is then sorted by seconds and the redundant observations are dropped. The data sets pl and nelson are then merged and the survival functions are plotted.
proc phreg data=datasets.honking noprint;
model seconds*censor(1) = ;
output out=pl logsurv=logs;
run;
proc sort data=pl;
by seconds;
run;
data pl;
set pl;
nlogs_pl = -logs;
by seconds;
if first.seconds;
run;
data combo;
merge nelson pl;
by seconds;
run;
goptions reset=all;
symbol1 c=blue i=stepjll;
symbol2 c=red i=stepjl line=21;
axis1 label=(a=90 'S(tj)') order=(0 to 3.5 by .5);
axis2 order=(0 to 20 by 5);
legend label=none value=(height=1 font=swiss 'Negative Log survivor' 'Nelson-Aalen' )
position=(top left inside) mode=share cborder=black;
proc gplot data=combo;
plot (nlogs_pl nlogs)*seconds / overlay vaxis=axis1 haxis=axis2 legend=legend;
run;
quit;