Version info: Code for this page was tested in R Under development (unstable) (2012-07-05 r59734)
On: 2012-08-08
With: knitr 0.6.3
Singular value decomposition (SVD) is a type of matrix factorization. For more details on SVD, the Wikipedia page is a good starting point. On this page, we provide four examples of data analysis using SVD in R.
Example 1: SVD to find a generalized inverse of a non-full-rank matrix
For a square matrix A with a non-zero determinant, there exists an inverse matrix B such that AB = I and BA = I. For a matrix that is not square, generalized inverse matrices have some (but not all) of the properties of an inverse matrix. SVD can be used to find a generalized inverse matrix. In the example below, we use SVD to find a generalized inverse B to the matrix A such that ABA = A. We compare our generalized inverse with the one generated by the ginv command.
library(MASS) a <- matrix(c(1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1), 9, 4) a.svd <- svd(a) a.svd$d
## [1] 3.46e+00 1.73e+00 1.73e+00 1.92e-16
ds <- diag(1/a.svd$d[1:3]) u <- a.svd$u v <- a.svd$v us <- as.matrix(u[, 1:3]) vs <- as.matrix(v[, 1:3]) (a.ginv <- vs %*% ds %*% t(us))
## [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] ## [1,] 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 ## [2,] 0.2500 0.2500 0.2500 -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 ## [3,] -0.0833 -0.0833 -0.0833 0.2500 0.2500 0.2500 -0.0833 -0.0833 ## [4,] -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 0.2500 0.2500 ## [,9] ## [1,] 0.0833 ## [2,] -0.0833 ## [3,] -0.0833 ## [4,] 0.2500
# using the function ginv defined in MASS ginv(a)
## [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] ## [1,] 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 0.0833 ## [2,] 0.2500 0.2500 0.2500 -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 ## [3,] -0.0833 -0.0833 -0.0833 0.2500 0.2500 0.2500 -0.0833 -0.0833 ## [4,] -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 -0.0833 0.2500 0.2500 ## [,9] ## [1,] 0.0833 ## [2,] -0.0833 ## [3,] -0.0833 ## [4,] 0.2500
Example 2: Image processing
The code below requires the ReadImages package. It reads in a jpeg (pansy.jpg) and plots it in R, first in color (when the image is stored as three matrices–one red, one green, one blue) and then in grayscale (when the image is stored as one matrix). Then, using SVD, we can essentially compress the image. Note that we can recover the image to varying degrees of detail as we recreate the image from different numbers of dimensions from our SVD matrices. You can see how many dimensions are needed before you have an image that cannot be differentiated from the original.
library(ReadImages) x <- read.jpeg("pansy.jpg") dim(x)
## [1] 600 465 3
plot(x, useRaster = TRUE)

r <- imagematrix(x, type = "grey") plot(r, useRaster = TRUE)

r.svd <- svd(r) d <- diag(r.svd$d) dim(d)
## [1] 465 465
u <- r.svd$u v <- r.svd$v plot(1:length(r.svd$d), r.svd$d)
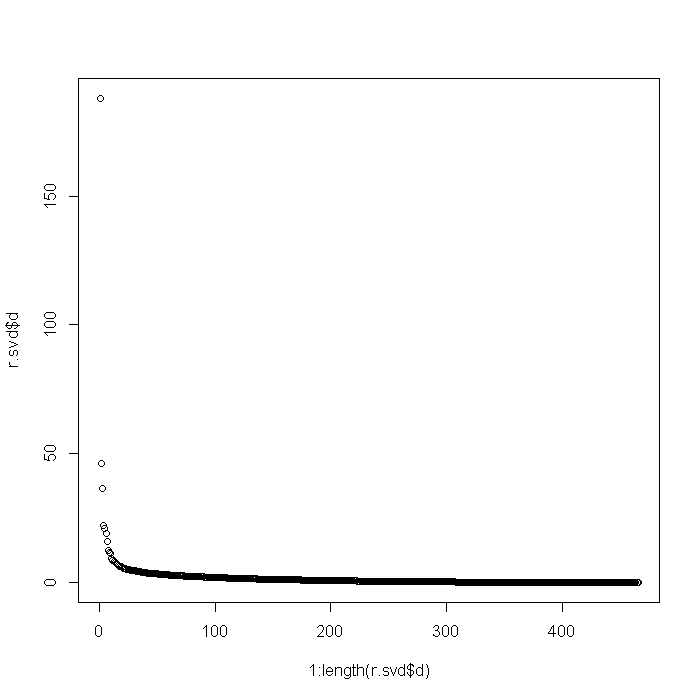
# first approximation u1 <- as.matrix(u[-1, 1]) v1 <- as.matrix(v[-1, 1]) d1 <- as.matrix(d[1, 1]) l1 <- u1 %*% d1 %*% t(v1) l1g <- imagematrix(l1, type = "grey") plot(l1g, useRaster = TRUE)
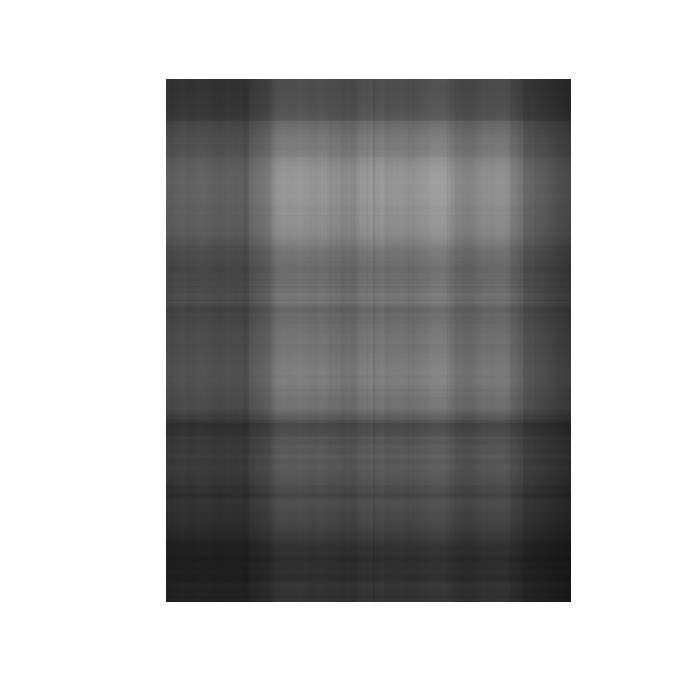
# more approximation depth <- 5 us <- as.matrix(u[, 1:depth]) vs <- as.matrix(v[, 1:depth]) ds <- as.matrix(d[1:depth, 1:depth]) ls <- us %*% ds %*% t(vs) lsg <- imagematrix(ls, type = "grey")
## Warning: Pixel values were automatically clipped because of range over.
plot(lsg, useRaster = TRUE)
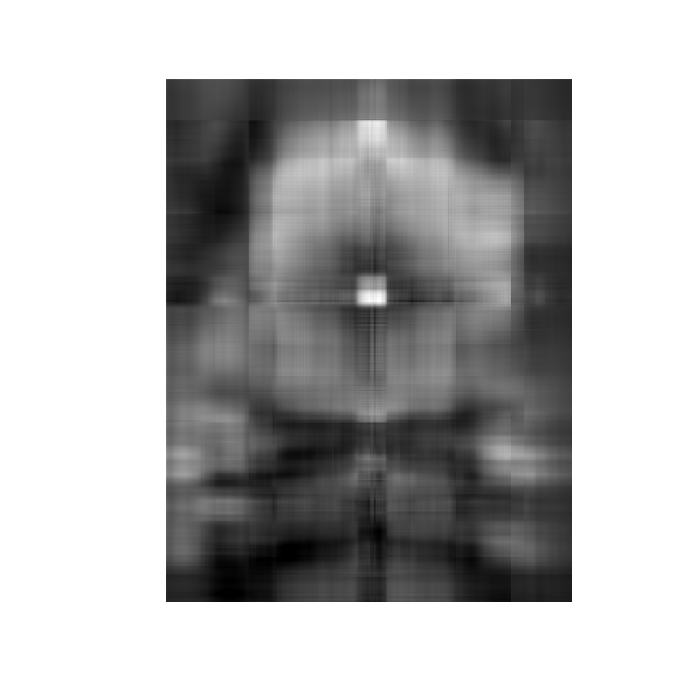

Example 3: Principal components analysis using SVD
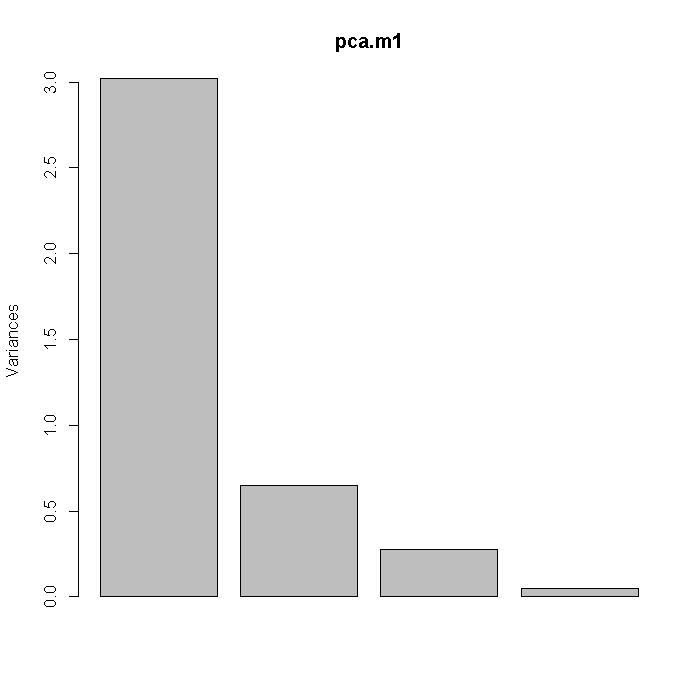
This example uses the Stata auto dataset. PCA can be achieved using SVD. Below, we first use the prcomp command in R and then plot the variances of the principal components (i.e. the square roots of the eigenvalues). These values can also be found through spectral decomposition on the correlation matrix or by SVD on the variable matrix after standardizing each variable.
library(foreign) auto <- read.dta("http://statistics.ats.ucla.edu/stat/data/auto.dta") pca.m1 <- prcomp(~trunk + weight + length + headroom, data = auto, scale = TRUE) screeplot(pca.m1)
# spectral decomposition: eigen values and eigen vectors xvars <- with(auto, cbind(trunk, weight, length, headroom)) corr <- cor(xvars) a <- eigen(corr) (std <- sqrt(a$values))
## [1] 1.738 0.807 0.526 0.225
(rotation <- a$vectors)
## [,1] [,2] [,3] [,4] ## [1,] -0.507 -0.233 0.825 0.0921 ## [2,] -0.522 0.454 -0.268 0.6708 ## [3,] -0.536 0.390 -0.137 -0.7358 ## [4,] -0.428 -0.767 -0.479 -0.0057
# svd approach df <- nrow(xvars) - 1 zvars <- scale(xvars) z.svd <- svd(zvars) z.svd$d/sqrt(df)
## [1] 1.738 0.807 0.526 0.225
z.svd$v
## [,1] [,2] [,3] [,4] ## [1,] 0.507 -0.233 0.825 -0.0921 ## [2,] 0.522 0.454 -0.268 -0.6708 ## [3,] 0.536 0.390 -0.137 0.7358 ## [4,] 0.428 -0.767 -0.479 0.0057
Example 4: Metric multi-dimensional scaling with SVD
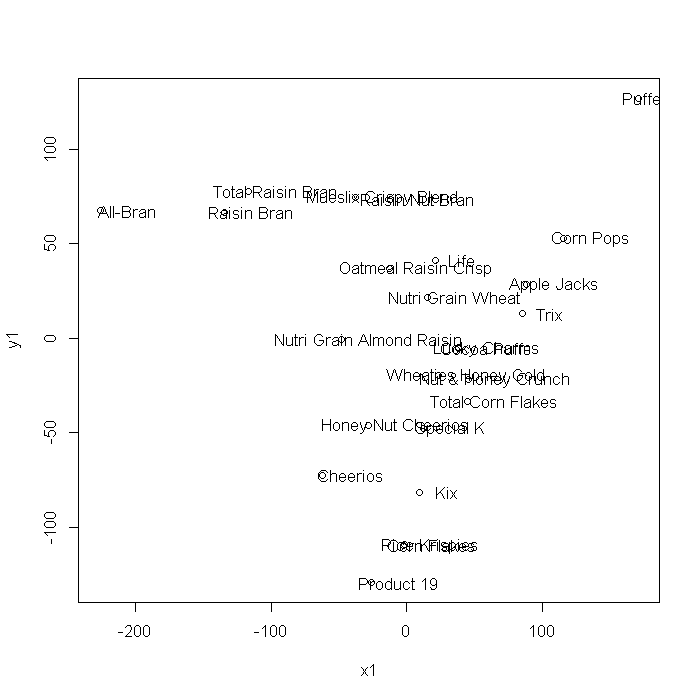
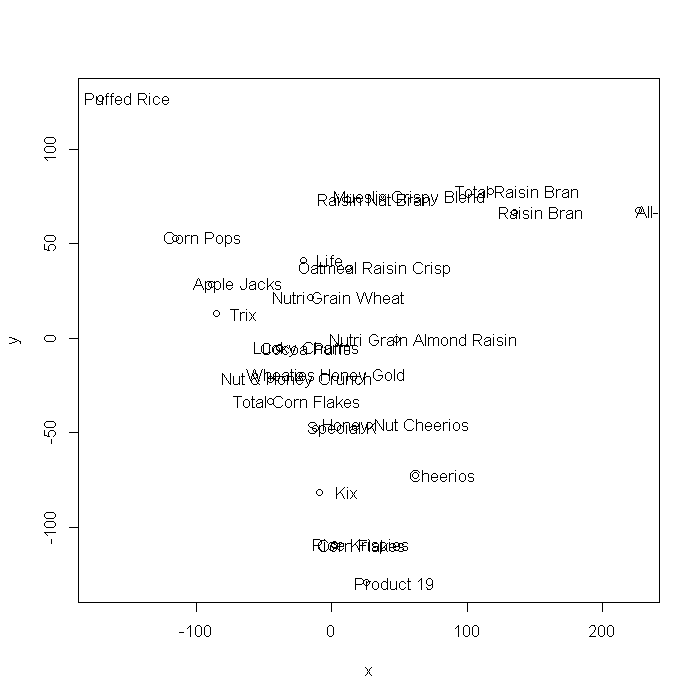
This example uses the Stata cerealnut dataset. Multi-dimensional scaling can also be achieved using SVD. The plots generated using cmdscale and the coordinates generated from the SVD steps are mirrored about the x/x1 = 0 axis, but are otherwise identical.
cnut <- read.dta("http://statistics.ats.ucla.edu/stat/data/cerealnut.dta") # centering the variables mds.data <- as.matrix(sweep(cnut[, -1], 2, colMeans(cnut[, -1]))) dismat <- dist(mds.data) mds.m1 <- cmdscale(dismat, k = 8, eig = TRUE) mds.m1$eig
## [1] 1.58e+05 1.09e+05 1.06e+04 3.83e+02 6.98e+01 1.25e+01 5.76e+00 ## [8] 2.22e+00 4.51e-12 4.51e-12 4.12e-12 3.19e-12 3.15e-12 2.30e-12 ## [15] 2.09e-12 1.25e-12 1.13e-12 8.79e-13 2.97e-13 -1.38e-12 -1.45e-12 ## [22] -1.57e-12 -1.88e-12 -5.92e-12 -2.52e-11
mds.m1 <- cmdscale(dismat, k = 2, eig = TRUE) x <- mds.m1$points[, 1] y <- mds.m1$points[, 2] plot(x, y) text(x + 20, y, label = cnut$brand)
# eigenvalues xx <- svd(mds.data %*% t(mds.data)) xx$d
## [1] 1.58e+05 1.09e+05 1.06e+04 3.83e+02 6.98e+01 1.25e+01 5.76e+00 ## [8] 2.22e+00 1.58e-11 9.90e-12 7.19e-12 4.71e-12 4.15e-12 3.03e-12 ## [15] 2.77e-12 2.08e-12 1.97e-12 1.50e-12 1.26e-12 1.05e-12 7.93e-13 ## [22] 7.35e-13 2.09e-13 1.65e-13 8.38e-14
# coordinates xxd <- xx$v %*% sqrt(diag(xx$d)) x1 <- xxd[, 1] y1 <- xxd[, 2] plot(x1, y1) text(x1 + 20, y1, label = cnut$brand)